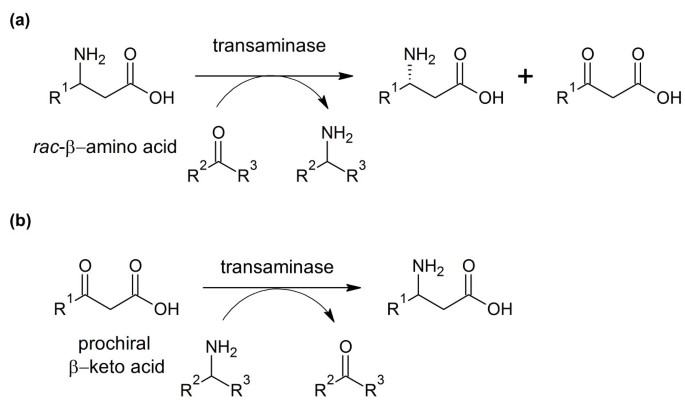
Catalysts | Free Full-Text | Recent Advances in ω-Transaminase-Mediated Biocatalysis for the Enantioselective Synthesis of Chiral Amines

Fluorescence-based high-throughput screening system for R-ω-transaminase engineering and its substrate scope extension | SpringerLink

PDF) Structural dynamics of the transaminase active site revealed by the crystal structure of a co-factor free omega-transaminase from Vibrio fluvialis JS17

Creation of (R)-Amine Transaminase Activity within an α-Amino Acid Transaminase Scaffold | ACS Chemical Biology

Structural dynamics of the transaminase active site revealed by the crystal structure of a co-factor free omega-transaminase from Vibrio fluvialis JS17 | Scientific Reports

Substrate profile of an ω-transaminase from Burkholderia vietnamiensis and its potential for the production of optically pure amines and unnatural amino acids - ScienceDirect

Structural dynamics of the transaminase active site revealed by the crystal structure of a co-factor free omega-transaminase from Vibrio fluvialis JS17 | Scientific Reports

RCSB PDB - 4A6T: Crystal structure of the omega transaminase from Chromobacterium violaceum in complex with PLP

PROCESS FOR THE IDENTIFICATION AND PREPARATION OF A (R)-SPECIFIC OMEGA- TRANSAMINASE - diagram, schematic, and image 44
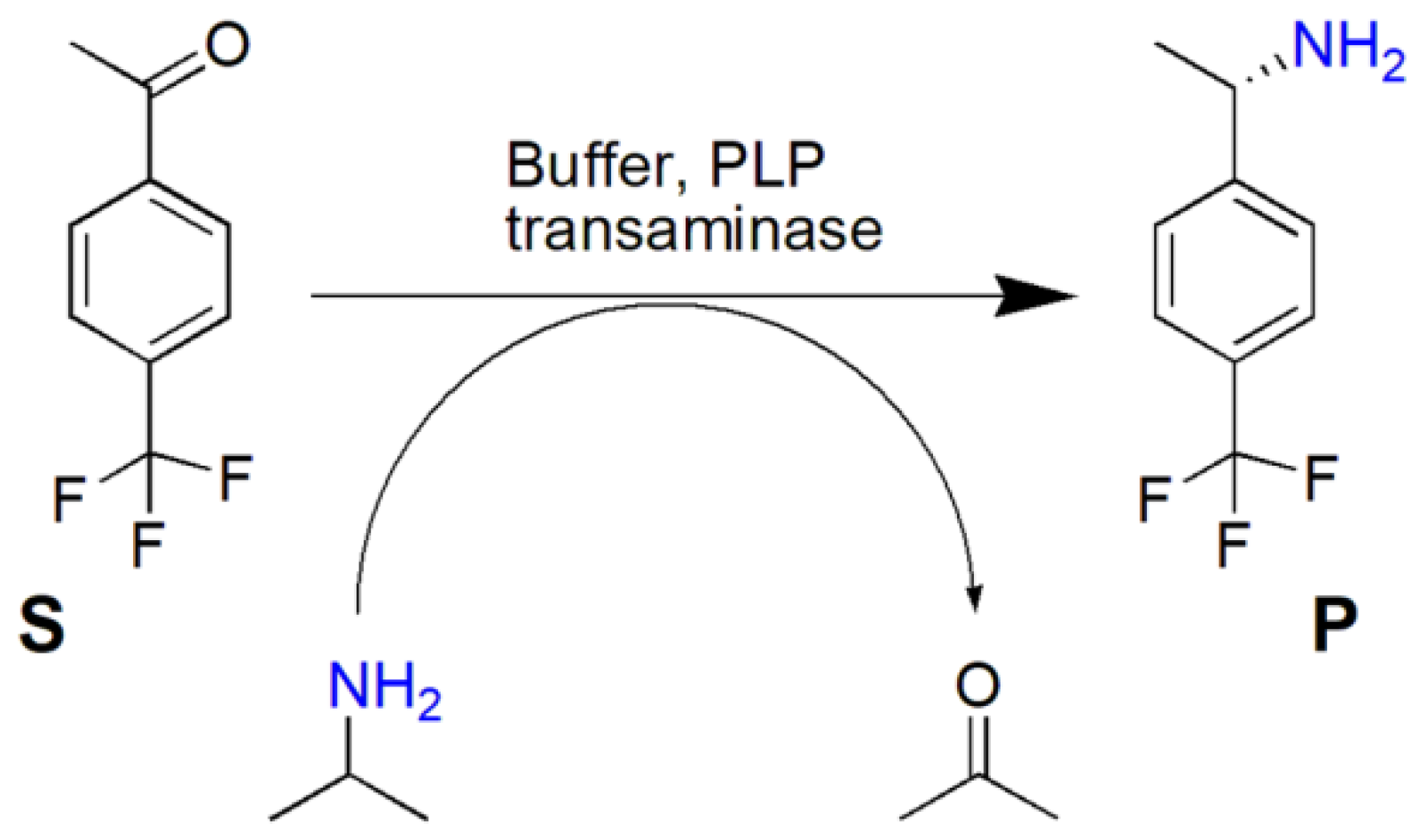
Catalysts | Free Full-Text | ω-Transaminase-Mediated Asymmetric Synthesis of (S)-1-(4-Trifluoromethylphenyl)Ethylamine
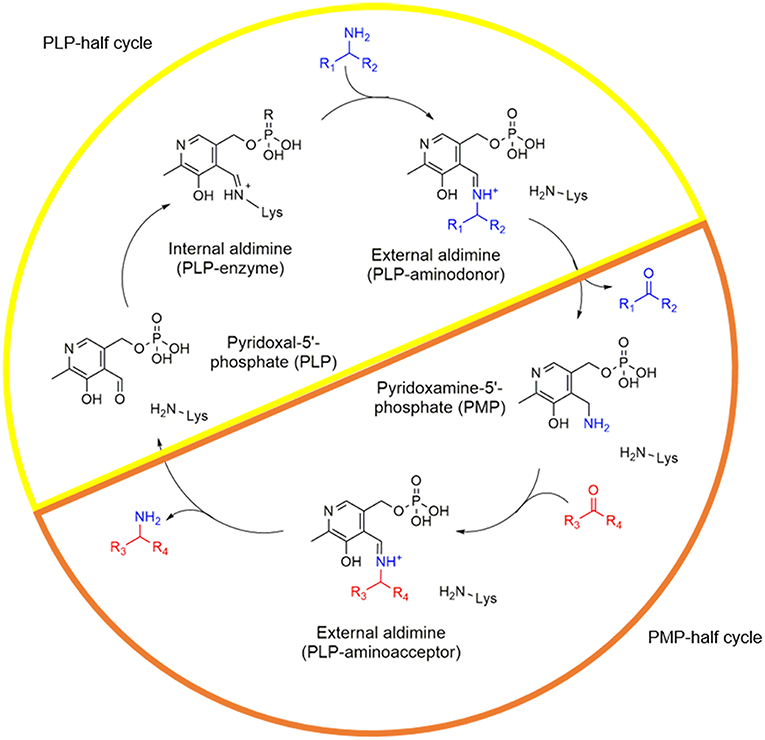
Frontiers | Enhancing PLP-Binding Capacity of Class-III ω-Transaminase by Single Residue Substitution

Mechanism‐Guided Computational Design of ω‐Transaminase by Reprograming of High‐Energy‐Barrier Steps - Yang - 2022 - Angewandte Chemie International Edition - Wiley Online Library

Computational Redesign of an ω-Transaminase from Pseudomonas jessenii for Asymmetric Synthesis of Enantiopure Bulky Amines | ACS Catalysis
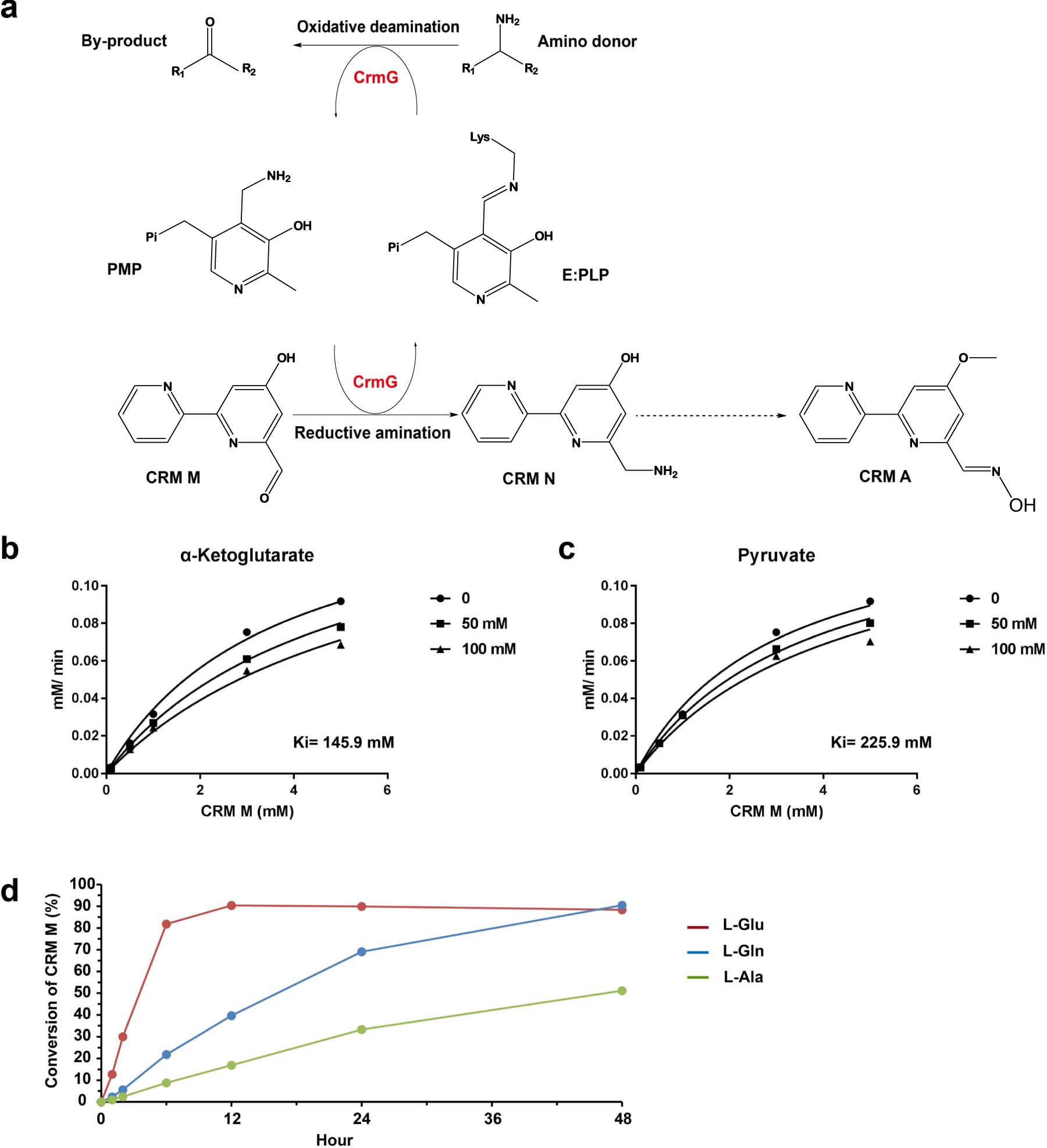
Structural studies reveal flexible roof of active site responsible for ω- transaminase CrmG overcoming by-product inhibition | Communications Biology

Redesign of (R)-Omega-Transaminase and Its Application for Synthesizing Amino Acids with Bulky Side Chain | SpringerLink

PROCESS FOR THE IDENTIFICATION AND PREPARATION OF A (R)-SPECIFIC OMEGA- TRANSAMINASE - diagram, schematic, and image 46